Handling of Missing Covariates
Meg Gonsalves
Postdoctoral Clinical Pharmacologist
Introduction
This blog post was led by Sravan Karumanchi, MS, Director of Pharmacometrics, and reflects a collaborative effort across our pharmacometrics team, with contributions from Kelly Larson, PhD, Taylor Eggertsen, PhD, Kun Yao, MA, Ethan DellaMaestra, BS, and Jonah Lyon, BS. Together they bring expertise in modeling, data strategy, and clinical interpretation.
In pharmacometrics (PMx), covariate handling is an essential component of both population pharmacokinetic (PopPK) and exposure-response (ER) analyses. In PopPK models, covariates help explain inter-individual variability in parameters such as clearance or volume, improving model interpretability and reducing unexplained variability. In ER analyses, covariates help identify subgroups that may differ in drug response or safety outcomes, and are often clinically meaningful for understanding benefit–risk across patient populations.
Covariates can be classified by their origin (intrinsic or extrinsic), measurement scale (continuous or categorical), and temporal nature (baseline or time-varying). Intrinsic covariates are subject-level variables such as demographics (e.g., age, weight, sex), physiological markers (e.g., renal or hepatic function), and measures of disease severity (e.g., tumor size, ECOG score), whereas extrinsic covariates are study-specific variables external to the subject (e.g., food effect or formulation). Continuous covariates span a range of numeric values (e.g., age, body weight, eGFR), while categorical covariates represent discrete groups (e.g., sex, race, renal impairment severity). Finally, baseline covariates are assumed constant from the time of first dose (e.g., sex, race), while time-varying covariates change over the course of the study (e.g., body weight, eGFR, tumor size). The collection of these potential covariates is meticulously scheduled during clinical trials and data is ultimately stored in CDISC standardized SDTM/ADaM domains such as DM (Demographics), VS (Vital Signs), LB (Laboratory Test Results). However, data collection is never perfect. Visits are skipped, tests are not collected, or results are populated as unknown resulting in missing information. Although data missingness is frequently inevitable, inappropriate handling of missingness could lead to biased results and incorrect conclusions which could have potentially devastating outcomes for patients. In a clinical setting, covariate misspecification may result in incorrect dose recommendations for a given population and inaccurate interpretations from exposure-response relationships.
There are several key components surrounding the handling of missing covariate data that will be outlined below.
Handling in PMx analyses
Data Assembly
Handling Missing Baseline Covariates:
Baseline covariates are often required for PMx analyses. While source data may explicitly flag the appropriate record as the study baseline, values can often be missing in source datasets due to operational or logistical reasons (e.g., labs collected after treatment started, screening failures, or missing CRF entries). A common strategy is to impute missing baseline values using the last non-missing observation prior to treatment initiation. These observations could be pulled from Screening or any other visits preceding the first dose date. Clinical judgement should be applied based on half-life of physiological stability to determine whether alternative time points can be substituted for baseline. For example, albumin changes slowly, so screening values are likely an accurate substitute for baseline, whereas vital signs are more variable and should have close temporal alignment whenever possible. If all imputation methods are exhausted, the variable should be documented as missing/unknown in the analysis dataset.
Handling Missing Time-varying Covariates:
Certain covariates may change over time during treatment. For time-varying analyses, it is important to ensure covariate values align temporally with the modeling data. A commonly used approach is to assign the most recent non-missing covariate value observed at or before that time, known as last observation carried forward (LOCF). Imputation with LOCF is appropriate for covariates that are relatively stable over time or change slowly, but may introduce bias for highly variable covariates or when time between observations is long.
Exploratory Data Analysis (EDA)
Categorical and continuous covariates with missing or unknown information should be evaluated for (1) extent of missingness and (2) contribution of effect and viability of combination. For categorical covariates, it may be appropriate to add an explicit "Missing" or "Unknown" category when there is substantial missingness or when imputation would have a confounding effect — for example, a race variable may combine "Missing" and "Other" categories if the concentration-time profiles of subjects in these groups appear similar. For continuous covariates, the potential effect of missingness on the analysis should similarly be assessed and reported. While no universal regulatory cutoff exists, published pharmacometric investigations commonly characterize <10% missingness as low and ≥50% as high, with decisions on handling missing covariates guided by the extent of missingness and its implications on the analysis.
Modeling
In model development, missing covariates will be imputed to covariate reference values. Thus, careful selection of covariate reference values are critical for pharmacometric analyses. Further, when a reference covariate is selected, the model parameters can be interpreted in the context of the reference subject. For example, if a reference weight of 70 kg and reference patient type of Non-Small Cell Lung Cancer (NSCLC) are chosen, the model clearance value will reflect that of the reference subject.
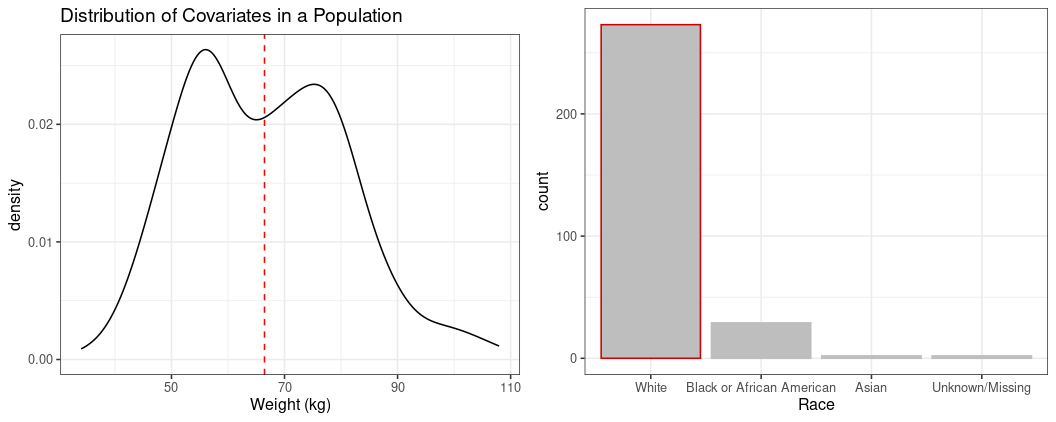
For continuous covariates, the median is frequently selected as the reference value, and for categorical covariates, the mode is frequently suggested as the reference value, as demonstrated in Figure 1.
When covariate missingness is substantial, or key covariates of interest are correlated, alternate analysis types may be considered beyond candidate covariate inclusion in model development. Such analyses may include investigating Empirical Bayes Estimates or post-hoc model predictions via categorizations of covariates that cannot be directly modeled. Additionally, sensitivity analyses may be performed to assess the influence of the highly missing covariate. Alternate analyses allow for exploratory investigation of covariate effects for variables with high missingness, while acknowledging the limitations caused by missing data.
Conclusion
Effective covariate handling is essential to ensure robust PMx analyses. Improper handling of missing covariates may lead to biased parameter estimates, false covariate relationships, masked true covariate effects, model instability or convergence errors, invalid predictions and simulation results, and loss of interpretability. To alleviate these concerns, decisions related to imputation, handling of missing data, and treatment of time-varying covariates should be grounded in biological plausibility, clinical relevance, and transparency. Clear documentation of data assumptions and imputation methods helps support model interpretability, reproducibility, and downstream communication with clinicians and regulatory stakeholders.